Originally published : Wed, February 11, 2015 @ 3:01 PM
Updated : Wed, March 29, 2023 @ 10:56 AM
As part of our “Know your oligo mod” blog series, we cover phosphorothioate bonds (PS), which are available for incorporation into custom oligonucleotides through LGC Biosearch Technologies.
This modification is interesting as it is not an addition of a chemical moiety per se, such as a nonstandard base or a quencher, but rather it is a special linkage between the bases. Specifically, it alters the standard phosphodiester linkages of DNA by replacing one of the non-bridging oxygen atoms with a sulphur atom, resulting in a phosphorothioate linkage (figure 1).
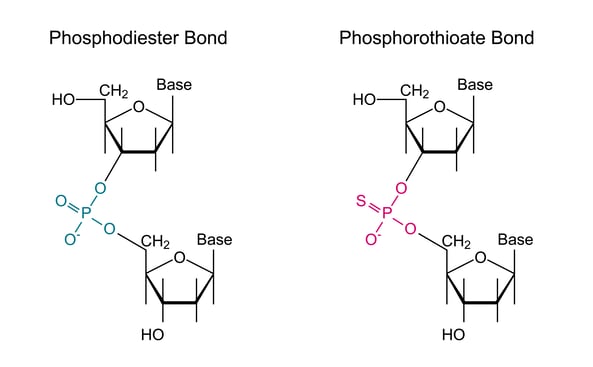 Figure 1: On the left, we see a schematic of a standard oligo linkage with a phosphodiester bond linking the two 5-carbon sugars through the phosphate bond forming the prototypical backbone of the oligo. On the right, we can see the phosphorothioate linkage in which a sulfur atom replaces one of the oxygens in the phosphate linkage between bases in the oligo.
Figure 1: On the left, we see a schematic of a standard oligo linkage with a phosphodiester bond linking the two 5-carbon sugars through the phosphate bond forming the prototypical backbone of the oligo. On the right, we can see the phosphorothioate linkage in which a sulfur atom replaces one of the oxygens in the phosphate linkage between bases in the oligo.
This alteration changes the overall chemical characteristics of the oligo, making it more suitable for various applications in molecular and cell biology research. It proves particularly valuable for therapeutic oligonucleotides like antisense oligonucleotides (ASOs), small interfering RNAs (siRNAs) and aptamers, enhancing their efficacy and utility in vivo.
What are the benefits of phosphorothioate bonds?
Nuclease resistance
Adding phosphorothioate bonds stabilises the oligo backbone against nuclease degradation, effectively enhancing the oligo’s half-life in the cellular milieu.1,2
This makes the modified bonds incredibly useful in the creation of antisense oligonucleotides, which when introduced into cells or biological matrices can interact with target nucleic acids to silence the expression of a particular transcript. Oligos containing phosphorothioate bonds accomplish this feat either through direct blockage of translation or enable enzymatic degradation of the target transcript, typically through an RNase H-mediated mechanism.3,4
Increased protein binding
Phosphorothioate moieties increase binding to various plasma and cellular proteins, which essentially determine the absorption and distribution of an oligo and are crucial for avoiding rapid clearance and enabling tissue entry.5,6
In vitro applications
PS-modified oligos aren’t just useful for therapeutic applications. They offer great protection wherever endo- and exonucleases may be found. They can be added to PCR primers to prevent degradation by DNA polymerases with proofreading activity,7 and are useful where the oligo may be transfected into a cell.
Enhancing the precision of CRISPR/Cas-mediated genome editing
Phosphorothioate modification has also been shown to be useful in CRISPR techniques. PS-modified oligos, in conjunction with an added bulky moiety, improved homology-directed repair (HDR) efficiency in a CRISPR/Cas genome editing study and prevented multiple templates from joining together before insertion. This allows for more controlled genome editing.8,9
Factors to consider when incorporating phosphorothioate modifications
Although phosphorothioate bonds confer superior properties, their repeated introduction can limit therapeutic function, induce toxic responses and affect oligo hybridisation kinetics. It is therefore important to realise that the phosphorothioate bond modification should not be freely incorporated at every available position in the oligonucleotide. Rather, they should be used sparingly with careful consideration of the impact on the overall functional properties. Here are some points to consider:
 Endo- or exonucleases
Endo- or exonucleases
For oligos requiring protection against exonucleases, it is recommended to incorporate the phosphorothioate bonds near the 5′ and 3′ ends of the oligo. However, for protection against endonucleases, these bonds can be included throughout the oligo sequence. In PCR, since polymerase extends from the 5' to 3', enhanced primers with PS bonds at both ends will result in a double-stranded product with PS bonds only at the 5' end. While protected from 5' to 3' nuclease degradation, these products lack PS bonds at the 3' end and will still be vulnerable to 3' to 5' nuclease degradation.
Melting temperature (Tm)
Increasing the number of phosphorothioate bonds in an oligo tends to lower the melting temperature (Tm) by around 0.5 °C per PS bond, thereby affecting hybridisation to the target.10,11
Reducing isomer effects
Phosphorothioate bonds introduced into the oligo create chiral centres at each bond, designated as either “Sp” or “Rp” conformation. This ultimately leads to multiple isomers of the oligo generated during synthesis.12 These individual isomers have been shown to have differential characteristics and functional properties. For example, the Rp isomer forms more stable duplexes with target RNA, but is more susceptible to exonuclease degradation, while endonucleases target the Sp isomer.13
Fortunately, much of the isomer effects on antisense oligos are mitigated through careful positioning of the modifications or by using additional modifications in conjunction with the phosphorothioate bonds; significantly diminishing concerns about non-racemic mixtures.14,15,16
GC content
Since PS-modified A:T pairs have been shown to have a greater reduction in stability than G:C pairs, it is suggested that GC-rich base sequences may be more suitable for PS-DNA oligos.17
Mitigating toxicity
The protein-binding ability of phosphorothioate bonds can have both positive and negative effects. While it enhances the in vivo and sub-cellular distribution of ASOs and siRNAs, it can also destabilise proteins, alter their cellular localisation, or lead to the formation of protein complexes, resulting in toxicities.18 As a consequence, the design* of therapeutic oligos can require significant trial and error to achieve the desired results.
Design and order custom oligonucleotides with phosphorothioate bonds
Biosearch Technologies offers synthesis of oligonucleotides incorporated with phosphorothioate bonds through our custom oligo service. Phosphorothioate bonds are universally represented with an asterisk (*) and designate the PS linkage when placed in between two bases. In the example “GTTA*CGTT” there will be a phosphorothioate bond in between the A and C base.
Pro-tip: We do not recommend Anion Exchange (AX) or DUAL (AX + RP) HPLC purification for oligos with more than a small number of phosphorothioates due to the potential for irreversible binding to the AX column. For oligos with more than a few phosphorothioate bonds, we recommend Reverse Phase (RP) HPLC.
Design and order custom oligos
Please note: Biosearch Technologies does not currently offer design services of antisense oligos or specific guidance in their use. We provide the custom synthesis and purification of these specialty modified oligos. If interested in additional information or resources about antisense oligonucleotides, please contact our technical support team.
Know your oligo mod series
- ATTO dyes
- 5-Nitroindole – a universal base oligo modification
- BHQ® (Black Hole Quencher®) non-fluorescent quenchers
- Methylene Blue
- Thio C6 linker
- 3' Spacer C3

|
Feeling ready to synthesise your own oligos?There are undeniable benefits to making your own oligos. However, beyond instrument selection there are a number of other considerations to work through such as solvent storage, disposal, ventilation, and other practicalities to ensure that your synthesizers run at full capacity and that you optimise your return on investment. Download a copy of this guide to support your decision making and a plan of action. |
References
- Vosberg HP, Eckstein F. Effect of deoxynucleoside phosphorothioates incorporated in DNA on cleavage by restriction enzymes. J Biol Chem. 1982;257(11):6595-6599.
- Crooke ST, Lebleu B. Antisense research and applications. CRC Press, 1993
- Kurreck J. Antisense technologies. Improvement through novel chemical modifications. European Journal of Biochemistry. 2003;270(8): 1628-1644. 10.1046/j.1432-doi:1033.2003.03555.x
- Stein CA. Phosphorothioate antisense oligodeoxynucleotides: questions of specificity. Trends Biotechnol. 1996;14(5):147-149. doi:10.1016/0167-7799(96)20006-X
- Crooke ST, Vickers TA, Liang XH. Phosphorothioate modified oligonucleotide-protein interactions. Nucleic Acids Res. 2020;48(10):5235-5253. doi:10.1093/nar/gkaa299
- Crooke ST, Seth PP, Vickers TA, Liang XH. The Interaction of Phosphorothioate-Containing RNA Targeted Drugs with Proteins Is a Critical Determinant of the Therapeutic Effects of These Agents. J Am Chem Soc. 2020;142(35):14754-14771. doi:10.1021/jacs.0c04928
- Skerra A. Phosphorothioate primers improve the amplification of DNA sequences by DNA polymerases with proofreading activity. Nucleic Acids Res. 1992. 25;20(14):3551-4. doi: 10.1093/nar/20.14.3551
- Gutierrez-Triana JA, Tavhelidse T, Thumberger T, Thomas I, Wittbrodt B, Kellner T, Anlas K, Tsingos E, Wittbrodt J. 2018. Efficient single-copy HDR by 5’ modified long dsDNA donors. eLife 7:e39468. 10.7554/eLife.39468
- Denes CE, Cole AJ, Aksoy YA, Li G, Neely GG, Hesselson D. Approaches to Enhance Precise CRISPR/Cas9-Mediated Genome Editing. Int J Mol Sci. 2021;22(16):8571. doi:10.3390/ijms22168571
- Uhlmann E, Peyman A, Will DW. Antisense: Medicinal Chemistry. Editor: Joseph R. Bertino. Encyclopedia of Cancer (Second Edition). Academic Press. 2002; Pages 103-117. doi:/10.1016/B0-12-227555-1/00012-5.
- Freier SM, Altmann K-H. The ups and downs of nucleic acid duplex stability: Structure-stability studies on chemically-modified DNA:RNA duplexes, Nucleic Acids Research, Volume 25, Issue 22, 1. 1997, 4429–4443, doi:10.1093/nar/25.22.4429
- Frey PA, Sammons RD. Bond order and charge localization in nucleoside phosphorothioates. Science 228.4699 (1985): 541-545. doi:10.1126/science.2984773
- Egli M, Manoharan M. Chemistry, structure and function of approved oligonucleotide therapeutics, Nucleic Acids Research, 2023;Volume 51, Issue 6, 2529–2573. doi:10.1093/nar/gkad067
- Koziołkiewicz M, Wójcik M, Kobylańska A, et al. Stability of stereoregular oligo(nucleoside phosphorothioate)s in human plasma: diastereoselectivity of plasma 3'-exonuclease. Antisense Nucleic Acid Drug Dev. 1997;7(1):43-48. doi:10.1089/oli.1.1997.7.43
- Koziolkiewicz M, Krakowiak A, Kwinkowski M, Boczkowska M, Stec WJ. Stereodifferentiation--the effect of P chirality of oligo(nucleoside phosphorothioates) on the activity of bacterial RNase H. Nucleic Acids Res. 1995;23(24):5000-5005. doi:10.1093/nar/23.24.5000
- Wan WB, Migawa MT, Vasquez G, et al. Synthesis, biophysical properties and biological activity of second generation antisense oligonucleotides containing chiral phosphorothioate linkages. Nucleic Acids Res. 2014;42(22):13456-13468. doi:10.1093/nar/gku1115
- Stein CA, Subasinghe C, Shinozuka K, Cohen JS. Physicochemical properties of phosphorothioate oligodeoxynucleotides. Nucleic Acids Research. 1988; 16(8),3209-3221. Doi: 10.1093/nar/16.8.3209
- Crooke ST, Seth PP, Vickers TA, Liang XH. The Interaction of Phosphorothioate-Containing RNA Targeted Drugs with Proteins Is a Critical Determinant of the Therapeutic Effects of These Agents. J Am Chem Soc. 2020;142(35):14754-14771. doi:10.1021/jacs.0c04928

