Originally published : Thu, May 24, 2018 @ 3:15 PM
Updated : Tue, May 29, 2018 @ 3:47 PM
Ever wake up in the morning just feeling like a rock star, or maybe like Francis Crick, Rosalind Franklin or Maurice Wilkins? You’re ready to just crush any task in front of you, discover something new and life changing. We get it. We’ve been there. And we love that we have products like Stellaris® RNA FISH – which is an incredibly straight forward technique that makes detecting, localizing and quantifying individual RNA molecules pretty simple. It’s times like this when you’re feeling invincible in your crisp white lab coat that crushing blows like forgetting to add a positive control to your Custom Stellaris probe set can derail your science even before it starts. It makes us cringe just thinking about it!
Don’t worry, you’re not alone, and hopefully with this simple checklist we’ve put together for you, some of these common pitfalls will be a thing of the past and you can get back to being a rock star,in and out of the lab of course.
I can see clearly now…
1. Make sure the dye you select is compatible with your microscope.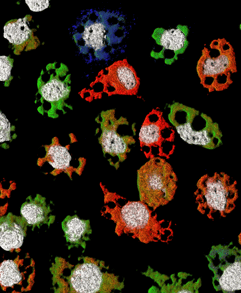 Our Stellaris Dyes and Modifications page provides important information on all the dyes available for Stellaris. You can check your dye filter set to see what fluorophores are compatible using our Chroma's spectra-viewer. This is a handy tool for checking the compatibility between your filter set(s) and the dye that you wish to use. Ideally, your microscope would have narrow filters that perfectly excites and captures the emission of our dyes.
Our Stellaris Dyes and Modifications page provides important information on all the dyes available for Stellaris. You can check your dye filter set to see what fluorophores are compatible using our Chroma's spectra-viewer. This is a handy tool for checking the compatibility between your filter set(s) and the dye that you wish to use. Ideally, your microscope would have narrow filters that perfectly excites and captures the emission of our dyes.
2. Choose a wide-field instead of confocal microscope
We recommend the use of wide-field fluorescence microscopes. Confocal microscopy uses point illumination to limit the focal plane for imaging. While this technique restricts light that is out of focus, it also diminishes the sensitivity of low-light level imaging. Some far-red dyes such as, Quasar 670® are susceptible to photobleaching caused by the intense laser light source from confocal microscopes.
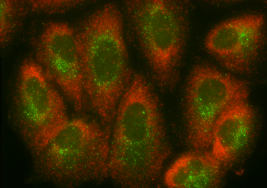 3. Do not select FAM as your designated dye for Stellaris RNA FISH
3. Do not select FAM as your designated dye for Stellaris RNA FISH
We advise against using the dye FAM due to the high cellular autofluorescence in that specific channel. If FAM must be used because there is no other choice, please ensure that the probe set labeled with this dye has a very high probe count, as close to 48 as possible, and that the application has been optimized to decrease autofluorescence. The FITC channel is generally easier to use in tissue culture than in tissue sections and should not be attempted in FFPE or high autofluorescence tissues, such as the brain.
Don’t be such a control freak…or actually...maybe…do.
1. Control your experiment with a ShipReady positive control probe set
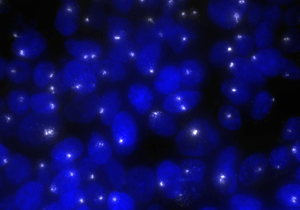 Positive controls are the backbone of every well-executed Stellaris experiment. Our ShipReady positive controls are essential for ensuring your Stellaris technique is sound and that your microscope is able to detect your selected dye. ShipReady probe sets recognize common genes, which are carefully selected reference genes, and long non-coding RNA targets with a predictable number of spots. Pre-verified ShipReady Probe Sets have been wet lab validated, which allows you to ensure that your Stellaris technique and microscope optics are sound before proceeding with custom designed probe sets.
Positive controls are the backbone of every well-executed Stellaris experiment. Our ShipReady positive controls are essential for ensuring your Stellaris technique is sound and that your microscope is able to detect your selected dye. ShipReady probe sets recognize common genes, which are carefully selected reference genes, and long non-coding RNA targets with a predictable number of spots. Pre-verified ShipReady Probe Sets have been wet lab validated, which allows you to ensure that your Stellaris technique and microscope optics are sound before proceeding with custom designed probe sets.
2. Run a negative, no probe control
By running a sample treated identically to your positive control, but without adding probes, you can observe what the natural autofluorescence of your tissue looks like. This step can help you to limit false-positive spot counting by recognizing what a negative result should look like.
3. Check to see if we offer a DesignReady RNA FISH probe set against your target gene
Check out our DesignReady targets to see which pre-designed assays we have available. DesignReady probe sets have been bioinformatically optimized by our professional design team.
4. Don’t see your target sequence as a DesignReady, you can design your own probe set.
If we don’t offer a probe set against your target RNA, you can design your own custom set using our online Stellaris probe designer software. Some parameters to consider are:
-Is your input sequence is at least 1,000 nucleotides long?
-Are at least 25 probes were generated with the online designer?- Stellaris probe sets with less than 25 probes in a set have a very low likelihood of success
-Did you read the helpful tips on our tech blog? The blog offers many other tips and design considerations to help design a custom probe set with the highest chance of success.
**Note: the success of a custom design probe is contingent on many parameters; ability to troubleshoot custom probe sets is limited without a purchased positive ShipReady control set
Stop, EVALUATE and listen
1. Do not perform Stellaris RNA FISH in FFPE tissue if you have any other option.
Using Stellaris RNA FISH for FFPE is not often successful as the technology requires high quality, intact RNA to allow for visualization of RNA. Most FFPE samples are preserved with protein in mind and not the more delicate RNA. This usually means longer times at higher temperatures and slower fixation or long lag times before harvesting the tissue, all of which degrade RNA. With that being said, we recommend using fresh-frozen tissue when given the option.
**if you decide to use FFPE, apply a ShipReady probe set alongside your custom or DesignReady probe set each time to confirm that there is intact RNA in the sample. It is essential to purchase a ShipReady to accompany your FFPE sample, as there is variation in RNA quality within a single block of tissue and even between regions in a single slice.
2. Label adherent cells first and tissue second.
First, validate your Stellaris RNA FISH experiment in adherent cells before moving on to using tissue samples. Tissue is known to have higher background autofluorescence, especially brain tissue, which is why you should validate this technique in adherent cells first.
3. Verify that your target is being expressed using qPCR
Before getting started with your Stellaris RNA FISH experiment, verify the expression of your target using qPCR. In some cases, your target might not be expressed in your experimental set up. It is always good to take precaution and to verify that your target is expressed.
There you have it rock star. While we know this list isn’t complete with hiccups that may arise, we hope that this checklist comes in handy when it comes time to light up your lab. And as always, if you have other issues let us know!

