Originally published : Fri, February 11, 2022 @ 5:57 PM
Updated : Fri, February 11, 2022 @ 10:58 PM
But measurement science could help steer us in a better, more informed direction
COVID-19 tests can indicate the presence of an infection, but not transmissibility
When a person contracts COVID-19, the amount of virus within the body, also known as the viral load or viral burden, changes as they progress from presymptomatic to symptomatic to recovery or continued disease. The amount of virus is what triggers a positive or negative result in a rapid antigen or PCR test, but how infectious that person may be over a period of time is still not well understood.1 Each type of test has its own limitations, but PCR is still considered the gold standard due to its sensitivity for detecting viral RNA. This test provides a quantitative output known as the quantification cycle (Cq), which is synonymous with the cycle threshold (Ct) or crossing point (Cp). While this value indicates the presence of viral RNA, it does not quantify the exact amount of virus within a patient nor does it accurately indicate infectivity.
Cq values are too variable to quantify virus in COVID-19
And yet, Cq cutoffs have been proposed to classify individuals at greater likelihood of spreading infection as well as performance targets for in vitro diagnostics. For example, the World Health Organization (WHO) published guidelines proposing that point-of-care tests need to detect a minimum Cq value of 25.2 As society craves for more definitive benchmarks to inform SARS-CoV-2 management such as quarantine periods, a recent study analysed Cq distributions of SARS-CoV-2 results from over 6,000 patients across three different clinical labs and found a 1,000-fold difference in copies/mL results for the same Cq value, which equates to approximately ten cycles.3
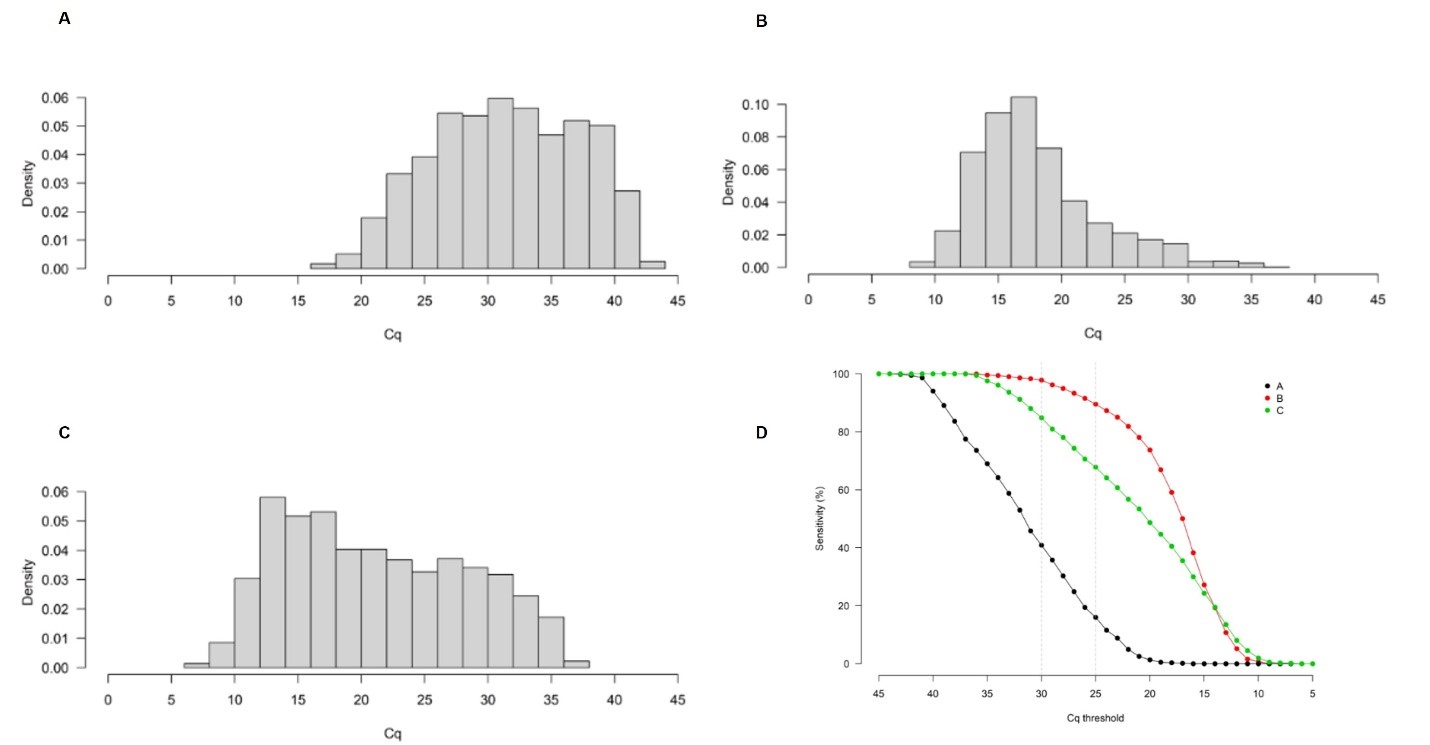
Figure 1. Histograms A, B and C show measured Cq values for three different cohorts within the study. Graph D depicts the impact of applying different Cq cutoffs on sensitivity. The plot shows for each cohort the proportion of positive samples which lie below a given cutoff. The vertical dotted lines indicate the sensitivity at Cq 25 and 30.
It is widely known that Cq values vary across labs due to several variables including nucleic acid extraction method, assay performance, platform and data analysis settings. The study revealed that after applying the Cq cutoff at 25 to the different cohorts, a clinical sensitivity of about 16% was observed in cohort A (United Kingdom) whereas it was about 90% in cohort B (Belgium). “This corresponds to a very large difference in viral copies per millilitre (copies/mL) measurement between one lab and another, despite the same Cq result,” Daniel Evans, the lead author of the study who is a PhD candidate in measurement science, notes, “our paper demonstrates that Cq as the main unit for patient stratification in COVID-19 produces highly variable results. Instead, using a copy-based unit of measurement would be relatively more accurate.” This can be achieved by using calibrated materials with qPCR to yield the underlying copy numbers corresponding to every Cq value.
Quantifying SARS-CoV-2 in copies/mL can better inform clinical decisions
Historically, viruses have been measured in international units or copy numbers, as seen in HIV, hepatitis B and hepatitis C. To evaluate the traditional measurement approaches used in viral quantitation, the three labs each generated copy number metrics using an external calibrator. These values were used to convert patient datasets into copies/mL. Although variability may still be introduced through the preanalytical factors like sample type and collection method, the inter-laboratory variability was reduced when comparing copies/mL. Jim Huggett, another author of the paper and Science Fellow at the UK National Measurement Laboratory at LGC, adds, “when a physician is informed that a patient has 10,000 copies of a virus versus 10 million copies, that makes it more clinically tangible than cycles 20 versus 30.” He recommends that clinical labs define how viral copy units correspond with their protocol’s Cq values.
Conducting a more strategic testing programme depends on the phase of the wave
Even with the use of calibrated copies/mL, the clinical sensitivity shifts depending on the viral distribution in the local population at the time of sample collection. For example, the viral load will be much higher in a region where the population is experiencing a surge in comparison to the lower viral load in another region that has a low positivity rate. A different study by Hay et al. measured SARS-CoV-2 viral RNA quantitative distribution within a population linked to its reproduction number (R). When R > 1, there are more individuals with newer infections, which at its peak can reflect a higher viral load in the population.4 Evans continues: “Upon looking at that data, we put forth a theory that for COVID-19 testing, less analytically sensitive methods, such as lateral flow tests, might be better suited during the peak of the pandemic when there is high incidence and the viral burden in the population is higher. But as cases drop, more sensitive methods, such as PCR tests, may be better to measure relatively lower levels of viral burden in the general population to avoid false negatives.”
Listen to a panel discussion corralling virological, diagnostic and clinical expertise on appropriate SARS-CoV-2 measurement: A lesson from COVID-19 on the dangers of using Cq to quantify nucleic acids.
References
- Walsh, K.A. et al. SARS-CoV-2 detection, viral load and infectivity over the course of an infection. J. Infect. 81(3): 357-371. Published June 29, 2020. Accessed February 4, 2022. https://dx.doi.org/10.1016%2Fj.jinf.2020.06.067
- World Health Organization. Target product profiles for priority diagnostics to support response to the COVID-19 pandemic v.1.0. Published September 28, 2020. Accessed February 1, 2022. https://www.who.int/publications/m/item/covid-19-target-product-profiles-for-priority-diagnostics-to-support-response-to-the-covid-19-pandemic-v.0.1
- Evans, D. et al. The Dangers of Using Cq to Quantify Nucleic Acid in Biological Samples: A Lesson From COVID-19. Clinical Chemistry. 86,(1): 153.162. Published October 22, 2021. Accessed February 1, 2022. https://doi.org/10.1093/clinchem/hvab219
- Hay, J.A. et al. Estimating epidemiologic dynamics from cross-sectional viral load distributions. Science. Vol 373. Issue 6552. Published June 3, 2021. Accessed February 4, 2022. https://doi.org/10.1126/science.abh0635


