Originally published : Tue, January 16, 2024 @ 1:56 PM
Since Humira (adalimumab) became the first approved monoclonal antibody therapy developed using phage display in 2002, it has sold more than any other drug in history.1,2 Phage display is a technique that allows for screening astronomical numbers of proteins for potent binders, and it is now one of the most powerful ways of creating high-affinity monoclonal antibodies in vitro.1
Phage display has been used for a range of purposes including developing diagnostic and therapeutic agents for transfusion medicine, autoimmune disease and neurological disorders.3 However, maximising the chance of success with this technique relies on selecting the most appropriate competent cells for transformation.
What's the appeal of phage display?
By inserting a gene coding for a protein of interest into the gene for a bacteriophage coat protein, Nobel laureate George Smith created an ingenious system to connect proteins with the gene that encodes them.4 This paved the way for efficiently screening enormous libraries of proteins and peptides based on their sequences.
The technique relies on a DNA vector that combines bacteriophage and plasmid properties, known as a phagemid. This allows infected E. coli competent cells to produce new phage particles complete with the protein or peptide of interest.
Phages that contain the modified gene “display” the protein of interest on their outer surface. With a library of different genes, the viruses can act as a platform for exposing billions of variations of a protein sequence to an immobilised target. The phages that stick to the target can be extracted and analysed to identify the best binders. This process, known as panning, can be repeated several times by amplifying the best binders from each round.
This screening process makes phage display particularly well-suited to discovering highly specific antibodies. It offers several advantages over hybridomas, an alternative approach to producing antibodies that fuses B cells with immortal cancer cells, including:
-
Covering a large diversity of clones
-
Screening toxic and non-immunogenic antigens
-
Making libraries from any animal, including humans
-
Having direct access to sequence, making it easier to further engineer antibodies
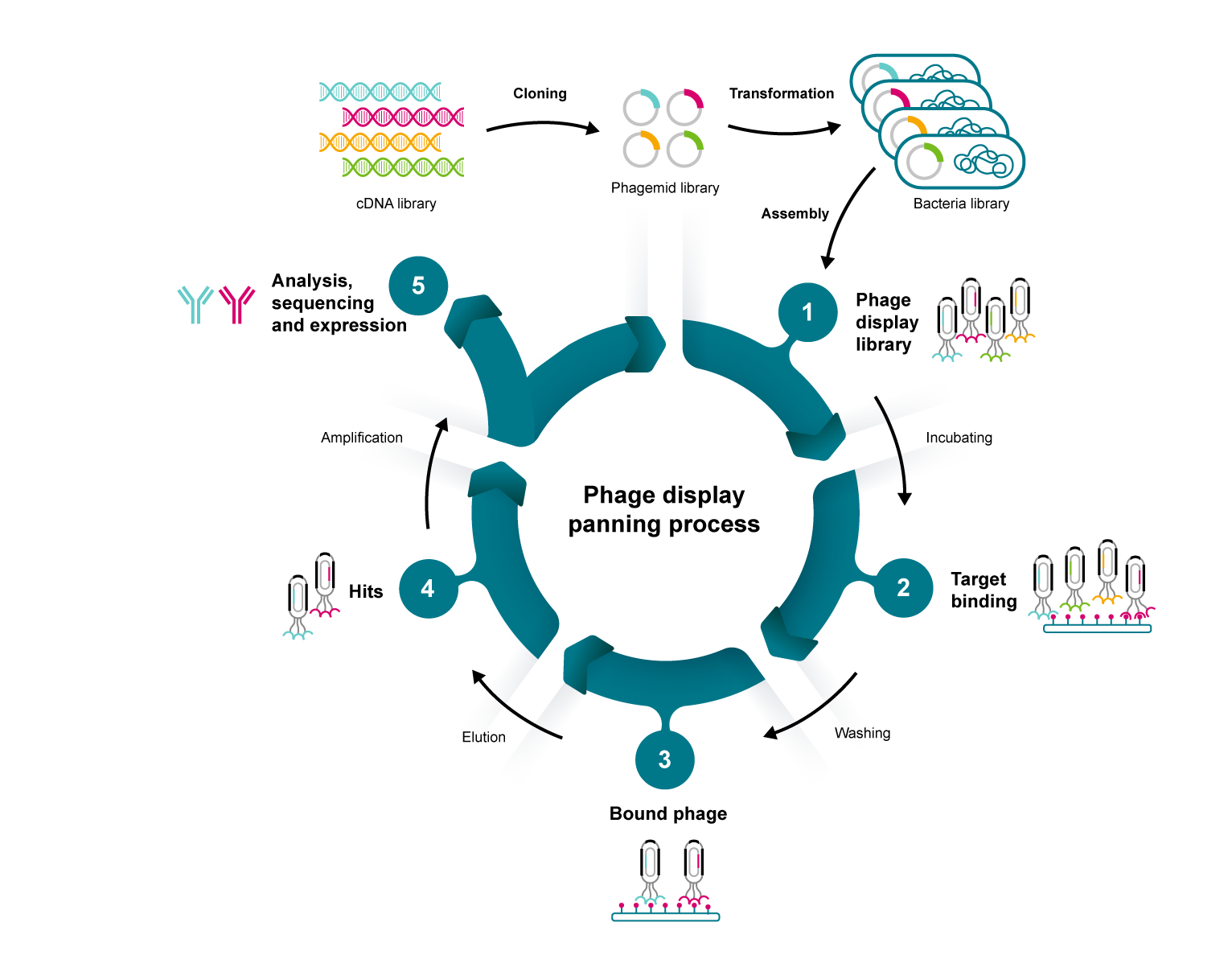 |
|
Figure 1. A representation of the panning process for phage display |
Why do competent cells matter?
When screening for the best possible binders, it is necessary to have a wide range of candidates to choose from. Phage display libraries can generate billions of different peptides and proteins.
|
|
To take advantage of this requires highly efficient competent cells. Greater transformation efficiency can dramatically increase the absolute number of recombinants for challenging applications, significantly reducing the cost to produce and screen phage display libraries of peptides and proteins. Cells optimised for DNA uptake can improve the transformation efficiency tenfold over existing methods.
Electrocompetent cells can achieve high transformation efficiencies, and therefore are recommended for producing large, high diversity libraries. These cells are engineered to have highly permeable membranes when electrical pulses are applied to allow the genetic material to easily penetrate.
Choosing the right type of competent cells
Several considerations are important for selecting the most suitable strain of competent cells for phage display:
-
As mentioned previously, the cells need to have very high transformation efficiency.
-
Users may want cells that are suitable for blue-white screening. This allows users to identify colonies carrying vectors with functioning inserts through their inability to express β-galactosidase and hydrolyse X-gal to produce a blue colour.
- Amber stop codons between the phage coat protein and the displayed peptide allow amber-suppressor strains to express the fusion protein.4
High-efficiency competent cells for a variety of applications
Biosearch Technologies offers a range of high-efficiency competent cells for different applications. Our most popular strains for phage display are:
-
TG1 – Ultra-high transformation efficiency, cloning methylated DNA, blue-white screening, amber suppressor strain, Fʹ plasmid. This is suitable for working with libraries of antibodies or antibody fragments.
-
SS320 (MC1061 Fʹ) – Ultra-high transformation efficiency, cloning methylated DNA, blue-white screening, Fʹ plasmid, non-amber suppressor.
-
ER2738 – High transformation efficiency, cloning methylated DNA, blue-white screening, amber suppressor strain, Fʹ plasmid. This is suitable for working with peptide display libraries.
-
MC1061F – Ultra-high transformation efficiency, cloning methylated DNA, blue-white screening. This is identical to strain SS320, except that it lacks the Fʹ episome that is required for infection by filamentous phage such as M13. The strain can be used for plasmid propagation and phage display but cannot be used for re-infection.
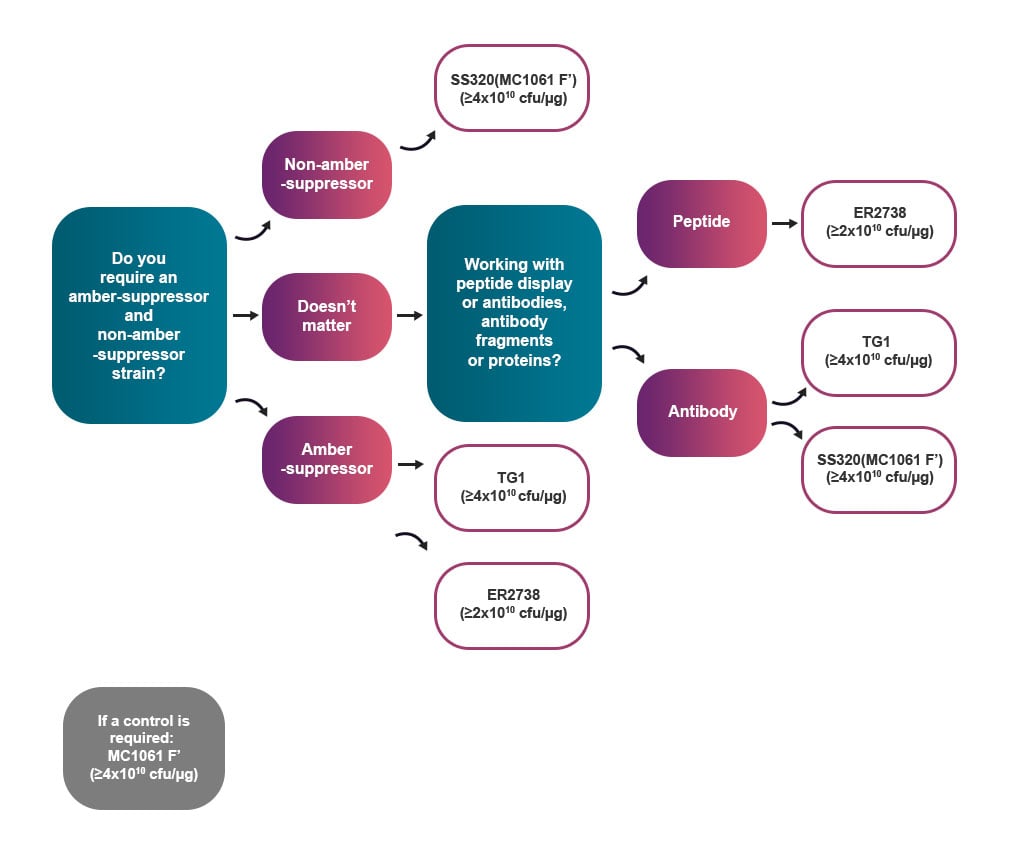
Select optimal cell strains for your experiment
The flowchart below can help you select the appropriate cell strain based on the type of phage display experiments that you are conducting.
From targeting tumours to treating respiratory diseases
Selecting the right competent cells is vital for achieving successful results. Our TG1 competent cells have been used in many high-profile studies to deliver ground-breaking phage display libraries, including from sharks, llamas and COVID-19 patients.
Researchers from the US National Cancer Institute developed a phage display library of 12 billion antibodies from nurse sharks.5 Cartilaginous fish, such as sharks, are the most ancient surviving animals that use antibodies in their adaptive immune system. The structure of shark antibodies allows them to be broken down into much smaller fragments than those based on human antibodies. This could help improve their tissue penetration, a key challenge in treating solid tumours.6
 The team reverse transcribed RNA extracted from the sharks’ blood, used PCR to amplify the sequences of interest, inserted them into a vector and used TG1 cells from Biosearch Technologies for transformation. The team showed that their library could identify binders against antigens associated with SARS, MERS, and liver and breast cancers.
The team reverse transcribed RNA extracted from the sharks’ blood, used PCR to amplify the sequences of interest, inserted them into a vector and used TG1 cells from Biosearch Technologies for transformation. The team showed that their library could identify binders against antigens associated with SARS, MERS, and liver and breast cancers.
This approach has also been done with miniature humanised antibodies derived from llamas.7 The library of 3 billion “nanobodies” could be used for specific protein knockdown in living cells and to target tumour-specific antigens.
Researchers have also used TG1 cells from Biosearch Technologies to create a library of patient-derived antibodies against SARS-CoV-2.8 This identified a panel of neutralising antibodies that bound to a range of positions on the virus, including some that remained conserved across different variants. This could allow the antibodies to target multiple circulating SARS-CoV-2 variants and those that might develop in future.
Taking care in your choice of cells
Success with phage display libraries relies on high efficiency competent cells to make the most of the power of this approach. The ability to make ground-breaking discoveries or world-leading drugs means that phage display is a technique that cannot afford to be neglected. By carefully choosing cells that are designed to suit your purpose, you can give your experiments the best chance possible of yielding the results needed.
For more information about how to select the best competent cells for your needs, check out our selection guide.
References
1. Nagano K and Tsutsumi Y. Phage Display Technology as a Powerful Platform for Antibody Drug Discovery. Viruses. 2021. 13(2): 178. doi: 10.3390/v13020178
2. Two decades and $200 billion: AbbVie’s Humira monopoly nears its end. Biopharma Dive. https://www.biopharmadive.com/news/humira-abbvie-biosimilar-competition-monopoly/620516/
3. Bazan J, Całkosiński I and Gamian A. Phage display—A powerful technique for immunotherapy. Human Vaccines and Immunotherapeutics. 2012. 8(12): 1817–1828. doi: 10.4161/hv.21703
4. Sioud M. Phage Display Libraries: From Binders to Targeted Drug Delivery and Human Therapeutics. Molecular Biotechnology. 2019. 61:286–303. doi: 10.1007/s12033-019-00156-8
5. Feng M et al. Construction and next-generation sequencing analysis of a large phage-displayed VNAR single-domain antibody library from six naïve nurse sharks. Antibody Therapeutics. 2019. 2(1):1–11. doi: 10.1093/abt/tby011
6. Nessler I et al. Increased Tumor Penetration of Single-Domain Antibody–Drug Conjugates Improves In Vivo Efficacy in Prostate Cancer Models. Cancer Research. 2020. 80 (6): 1268–1278. doi: 10.1158/0008-5472.CAN-19-2295
7. Moutel S et al. NaLi-H1: A universal synthetic library of humanized nanobodies providing highly functional antibodies and intrabodies. eLife. 2016. 5:e16228. doi: 10.7554/eLife.16228
8. Bullen G et al. Cross-Reactive SARS-CoV-2 Neutralizing Antibodies From Deep Mining of Early Patient Responses. Frontiers in Immunology. 2021. doi: 10.3389/fimmu.2021.678570