Originally published : Thu, October 30, 2014 @ 6:27 PM
Updated : Wed, March 22, 2023 @ 1:23 PM
Stellaris controls, who needs ‘em? We all do. Especially if it’s your first time interrogating a sample with a new Stellaris RNA FISH probe set, it is critical to consider the proper controls. The concept of positive and negative controls for experiments is not a new one. Far too often, we tend to bypass the consideration of controls in favour of saving time. However, it really all boils down to, “how do you, as a scientist, interpret your results after the experiment is over?”
Positive and negative controls for Stellaris experiments can be viewed from multiple angles. Let’s say you were successful in measuring spots on your first try, the major questions you may ask are:
- Is the signal I am measuring being produced by probe bound to RNA?
- Are these spots true signal or just background?
- Are these spots specific to my target or are they picking up other RNAs?
But let’s say the opposite happened, you didn’t get any spots at all. You may ask yourself:
- Did I perform the experiment correctly?
- Is my RNA expressed in my sample or is it degraded?
Below, are some recommendations for controls when performing a Stellaris experiment. This is not an exhaustive list of potential controls, and you may find ways to better interpret your results through alternate controls. If so, feel free to share them in the comments section.
1. Is the signal I am measuring produced by probe bound to RNA?
Suggested control: Treat with RNase
This particular question may be asked the first time you perform a Stellaris experiment with a new probe set. Likely, you will only need to include this control once in order to convince yourself (or a reviewer!) that the signal you are picking up is indeed due to RNA and not some other extraneous signal. A simple way to answer this question is by using an RNase, pre-treated control sample. Usually, this involves treating a sample with RNase A (50 µg/mL) for 30 min. to 1 hr at 37 °C, prior to the hybridization step. There are other variations to this general protocol and you may have to adjust as necessary for your sample type. If your Stellaris signal is produced by probe bound to RNA, then the signal should disappear with RNase treatment. <<Click to Tweet>>
2. Are these spots true signal or just background?
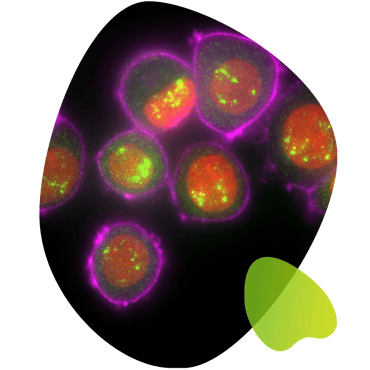 You have finished your experiment and you are excited that you got a signal. Congratulations! But doubt may soon rear its ugly head...what if these spots aren’t real? In order to avoid these doubts, it is imperative to include a number of controls for signal specificity.
You have finished your experiment and you are excited that you got a signal. Congratulations! But doubt may soon rear its ugly head...what if these spots aren’t real? In order to avoid these doubts, it is imperative to include a number of controls for signal specificity.
Suggested control: No-probe control
The first control to consider is a no-probe control sample. This sample should be treated the same as the rest of your samples, but probed with hybridization buffer only. This no-probe control sample will allow you to distinguish true spots from autofluorescence.
Suggested control: Image All Samples in an unused filter
When imaging all of your samples, it will also be critical to image in an unused filter (one that your dye should not be detected in) to determine non-specific signal. A FITC filter has become a standard filter block in almost every microscopy brand and platform. A typical FITC excitation filter block is 475-495 nanometers and the emission is 500-540 nanometers. This is usually our go-to choice for measuring autofluorescence in any sample, specifically if you are working with tissue. Keep in mind that autofluorescence can be highly variable in shape and size. If these same spots or shapes appear in an unused filter, then they are certainly suspect.
Suggested control: Target a gene from an unrelated organism
In addition to the above recommendations, you may consider using a probe set targeting a gene from an unrelated organism. For example, if your cells or tissue do not express GFP (green fluorescent protein), a probe set targeting GFP RNA can be used. Customers often ask if they should use sense probes as negative controls. We generally don’t recommend this type of approach for the Stellaris technology because this may just lead to higher background, false signal, or the very real possibility of transcription from the sense strand!
3. Are these spots specific to my target or are they picking up other RNAs?
We suggest: Use a cell line or tissue void of the target transcript
To determine the specificity of your signal from the gene-specific probe set, an ideal negative control is to test the probe set in a cell line or tissue void of the transcript. Also possible is to use cells or tissue with a targeted siRNA knockdown or knockout, respectively. As long as care has been taken to design your probe set using a specific RNA sequence, this usually is not a problem. However, there are many RNAs with splice variants, or RNAs that come from large gene families with highly similar sequences. Careful probe set design is key. We have design tips and recommendations available in our Guide to Getting Started with Stellaris if you would like more information on this very important topic.
4. Did I perform the experiment correctly?
Often, you ask this question when you get no result at all. It’s a hard question to answer when you don’t have a positive control. Also, there may be no result when the RNA integrity of your sample is suspect, or if the RNA you are targeting is not actually expressed in your sample. There are a number of ways to avoid having to ask this question after you have finished your experiment.
We suggest: Use a catalogued probe set
Try one of our catalogued, functionally tested positive control probe sets when performing your experiments. If the experiment is performed properly and the positive control gene is expressed in your sample type, a specific signal will be produced. If using a catalogued probe set is not an option, design a custom probe set to a gene target that has relatively medium to high abundance in your sample type in addition to designing a custom probe set to your target gene.
5. Is my RNA present or too low for detection in my sample or is it degraded?
Suggested control: Perform a complementary method
When you are testing a sample for RNA expression the first time, you will want to ensure that your target is actually expressed in your sample type by running a complementary method, such as qPCR. Performing qPCR will also help give you an idea of the abundance of your target in your sample.
Suggested control: Over-express the target in a cell line if possible
In some cases, the RNA signal may be difficult to distinguish, possibly because of low copy number. In this case, you may consider increasing the expression your target by transfecting or transducing the target into a cell line in order to confirm the specificity of your signal. If your target can be induced with a drug or some other treatment, this may also be a good avenue to explore.
Suggested control: Check integrity of RNA in sample
It will also be important to check the integrity of the RNA in your sample. There are a number of ways to do this in addition to using a positive control probe set (as described above).
Do you have more questions for our technical team? Leave a comment below or submit your query here.

