Originally published : Fri, January 21, 2022 @ 10:30 PM
Updated : Tue, January 25, 2022 @ 8:30 PM
More innovative applications for Black Hole Quencher (BHQ) dyes
In imaging and detection assay development, innovative approaches are often needed to push the boundaries of what we can uncover. LGC, Biosearch Technologies™ is proud to enable many of these advances with our family of Black Hole Quencher™ (BHQ™) dyes. In this second blog post of a two-part series, we highlight more exciting imaging and detection applications that leverage the power of BHQ dyes.
Read the first blog post here, and learn more about the advantages of BHQ dye technology here.
BHQ dyes enable simultaneous in vivo monitoring of insulin and glucose
The challenge
Understanding how our internal biochemistry changes over time and responds to therapeutic interventions can enhance human health, enable precision medicine and accelerate therapeutic development. But the development of sensors that can monitor multiple analytes in real time at physiologically-relevant concentrations has been challenging and limited to only a few analytes, such as glucose.
The solution
In a recent paper, Poudineh et al.1 report the development of a microfluidic system that can simultaneously monitor glucose and insulin in real time in a diabetic rat model, and is sensitive enough to detect and differentiate between the pharmacokinetics of short-acting versus delayed-onset insulin formulations.
Part of what makes their system so exciting is that it is easily generalisable to detect a wide range of analytes.
The technological advances
To achieve this significant development, Poudineh et al.1 combined a specially-designed microfluidic chip with a bead-based sandwich ELISA assay to measure insulin and a bead-based aptamer assay to measure glucose.
How BHQ dyes helped
BHQ dyes were used to keep background low during glucose measurement. The authors attached glucose-recognising aptamers labelled with a Cy5 fluorophore to beads, and used complementary oligos with a BHQ dye to quench Cy5 signal in the absence of glucose. In the presence of glucose, the BHQ oligo gets displaced and the Cy5 signal can be detected (Figure 1). The fluorescent signal intensity of the beads was shown to increase with concentration across a physiologically-relevant range, enabling quantification of the analyte.
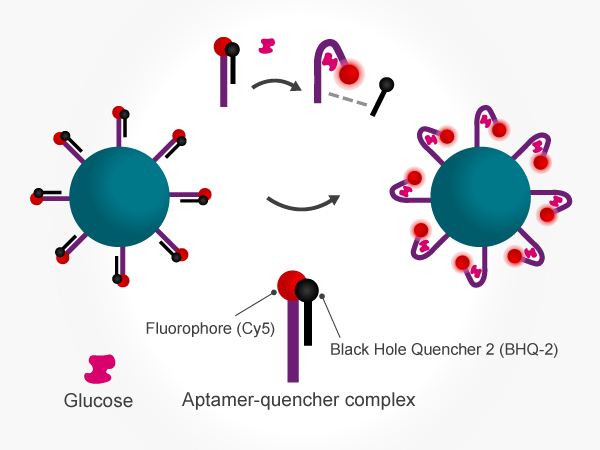 Figure 1. Continuous measurement of analytes using a bead-based fluorescence assay. Glucose aptamers labelled with Cy5 fluorophore were attached to beads. Hybridised complementary DNA strands conjugated to a BHQ-2 dye were displaced in the presence of glucose, eliminating quenching and producing a fluorescent signal.
Figure 1. Continuous measurement of analytes using a bead-based fluorescence assay. Glucose aptamers labelled with Cy5 fluorophore were attached to beads. Hybridised complementary DNA strands conjugated to a BHQ-2 dye were displaced in the presence of glucose, eliminating quenching and producing a fluorescent signal.
BHQ dyes drive detection of signal pathway activation rather than individual signal transduction proteins
The challenge
Signaling pathways are critical for many cellular processes, but pathways are often complex and overlapping, with individual protein components shared across multiple distinct pathways. Because of this overlap, tracking activation of a specific signaling pathway often requires monitoring multiple components, making studies complicated and labour-intensive.
The solution
Shi et al.2 provide an elegant solution to this challenge by developing a detection system that only delivers a signal when both an upstream and a downstream component of the signaling pathway are present. In this proof-of-concept paper, they demonstrate the potential of their sensor to detect activation of an apoptotic signaling pathway that travels through manganese superoxide dismutase (MnSOD) and not when the apoptotic signal travels through c-Myc, as well as the reverse – a signal that travels through c-Myc and not through MnSOD.
The technological advances
To create a sensor that only signals when two components are present – what the authors refer to as a logic gate performing an “and” operation – they created a Y-shaped DNA that acts as the sensor (Figure 2). One branch of the Y contained a complementary sequence for binding MnSOD mRNA, the upstream pathway molecule and the tether to a gold nanoparticle. The other branch of the Y contained an aptamer for cytochrome c (Cyt c), the downstream pathway molecule, and a Cy5 fluorophore. Lastly, a DNA strand containing a BHQ dye spanned the two branches of the Y together.
Binding of the MnSOD mRNA to the sensor releases the BHQ-containing DNA and aptamer from the gold nanoparticle, but the presence of the BHQ dye prevents Cy5 fluorescence. Only when Cyt c binds to the aptamer is the BHQ dye-containing DNA released, resulting in a fluorescent signal.
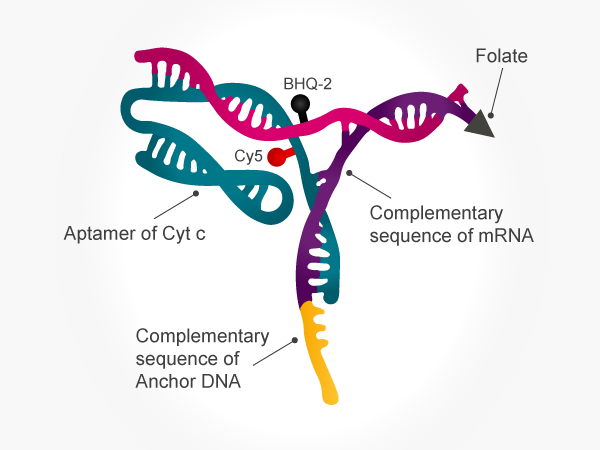 Figure 2. Two-component sensor developed by Shi et al.2 Yellow indicates the tether to the gold nanoparticle; purple indicates the MnSOD mRNA binding domain; teal indicates the Cyt c aptamer; pink indicates the spanning DNA with the BHQ dye; red indicates the Cy5 fluorophore. The folate facilitates entry into cells by receptor-mediated endocytosis.
Figure 2. Two-component sensor developed by Shi et al.2 Yellow indicates the tether to the gold nanoparticle; purple indicates the MnSOD mRNA binding domain; teal indicates the Cyt c aptamer; pink indicates the spanning DNA with the BHQ dye; red indicates the Cy5 fluorophore. The folate facilitates entry into cells by receptor-mediated endocytosis.
How BHQ dyes helped
The use of the BHQ dye was critical for keeping fluorescence quenched until both molecules, MnSOD and Cyt c, were detected.
BHQ dyes support sensitive detection of sepsis-causing bacteria from blood
The challenge
Sepsis can be potentially fatal and is often mistaken for other pathologies, leading to a delay in appropriate life-saving treatment. The challenge lies in being able to rapidly detect bacterial pathogens, which can often be present in blood at very low concentrations. Current diagnostic methods can take several days while the potential bacteria are cultured, during which time the disease can worsen.
The solution and technological advances
Fang et al.3 developed an integrated microfluidic chip that uses membrane-based filtration to separate red and white blood cells from the bacteria as well as mannose-binding lectin-coated beads to capture both Gram positive and Gram negative bacteria. Bacteria were identified using on-chip hydrolysis probe qPCR technology.
How BHQ dyes helped
An integral part of hydrolysis probe technology, BHQ dyes are a best in class quencher, driving sensitive detection using qPCR.
BHQ dyes support accurate detection and quantification of a genome edited crop
The challenge
Some countries regulate genome edited crops in the same way as genetically modified organisms (GMOs), requiring that they be detected and distinguished from naturally occurring variants. The challenge is that while conventional GMOs typically introduce foreign DNA, the presence of which clearly indicates that the crop is a GMO, genome edited crops do not.
The solution
Zhang et al.4 noted that their lab had used genome editing on rice to add or delete 1-18 base pairs in the chlorophyllide a oxygenase 1 (CAO1) gene. They created a series of primer and probe sets designed to specifically detect and quantify these insertions and deletions using both qPCR or ddPCR.
The technological advance
To increase the specificity of detection and reduce oligo annealing to off-target sequences, they used locked nucleic acid probes which provide increased stability at higher temperatures compared to the conventional probe.
How BHQ dyes helped
As always, BHQ dyes are essential for sensitive detection and quantification using qPCR and ddPCR.
How will BHQ dyes power your innovation?
As you can see from these published examples, BHQ dyes enable a wide range of innovative applications. We look forward to helping you push the frontiers of detection and imaging with BHQ dyes, and hope to be able to discuss your exciting advance in a future blog post!
References
- Poudineh M, Maikawa CL, Ma EY, Pan J, Mamerow D, Hang Y, Baker SW, Beirami A, Yoshikawa A, Eisenstein M, Kim S, Vučković J, Appel EA, Tom Soh H. A fluorescence sandwich immunoassay for the real-time continuous detection of glucose and insulin in live animals. Nature Biomedical Engineering. 5,(1) 53–63. https://doi.org/10.1038/s41551-020-00661-1. Published 2020. Accessed January 17, 2022.
- Shi H, Wang Y, Zheng J, Ning L, Huang Y, Sheng A, Chen T, Xiang Y, Zhu X, Li G. Dual-Responsive DNA Nanodevice for the Available Imaging of an Apoptotic Signaling Pathway in Situ. ACS Nano. 26;13(11):12840-12850. https://doi.org/10.1021/acsnano.9b05082. Published 2019. Accessed January 17, 2022.
- Fang YL, Wang CH, Chen YS, Chien CC, Kuo FC, You HL, Lee MS, Lee GB. An integrated microfluidic system for early detection of sepsis-inducing bacteria. Lab Chip. 21, 113-121. https://doi.org/10.1039/D0LC00966K. Published 2021. Accessed January 17, 2022.
- Zhang H, Li J, Zhao S, Yan X, Si N, Gao H, Li Y, Zhai S, Xiao F, Wu G, Wu Y. An Editing-Site-Specific PCR Method for Detection and Quantification of CAO1-Edited Rice. Foods.10(6), 1209. https://doi.org/10.3390/foods10061209. Published 2021. Accessed January 17, 2022.


