Originally published : Tue, August 4, 2015 @ 3:49 PM
Updated : Wed, March 29, 2023 @ 10:38 AM
Chemically synthesized DNA has become an indispensable tool for molecular biologists in the fields of medicine, diagnostics, agriculture, and biological research. Today, chemically synthesized DNA in the form of probes, primers and other oligonucleotides are used in a variety of applications such as PCR, genome sequencing, SNP genotyping, and gene synthesis. While oligonucleotides, or oligos, have become commonplace in these applications, they should not be viewed as a commodity. Rather, they represent critical components providing the molecular mechanism for a biochemical event.
In nature, DNA is assembled enzymatically in the 5’ to 3’ direction. In contrast, the manufacture of oligos is now done through chemical synthesis in the 3’ to 5’ direction. However, this was not always the case. As nucleic acid chemistry evolved, the direction of the synthesis also evolved. It has taken years of research and investigation to advance small and large scale oligo synthesis so that their manufacture is now routine.
This article will highlight some of the prominent and dedicated scientists who developed nucleic acid chemistry, and present recent advancements in the manufacture of probes or primers as synthesized today.
Historical DNA synthesis chemistry
The first dinucleotide was chemically synthesized in the 5’ to 3’ direction using solution phase chemistry. In 1955, Michelson and Todd reported the preparation of a dithymidinyl nucleotide1. This was the first published paper in which two deoxythymidine bases were synthetically prepared and reacted by chemical means to form a dinucleotide. The chemical synthesis of DNA proceeded using this methodology until the 1960’s.
In the late 1950’s, scientist H. Gobind Khorana invented the phosphodiester method of oligo synthesis and would later share the Nobel Prize for investigation into the genetic code 2. The phosphodiester method is a solution phase chemistry which introduces a stable nucleoside phosphine that will couple to a desired nucleotide when activated with DCC (N,N’-dicyclohexylcarbodiimide) 3. Through this method , Khorana and his team were able to chemically synthesize a much longer oligo, in the 5’ to 3’ direction, corresponding to the sequence of a transfer RNA 4.In Khorana’s quest to interpret the genetic code and its function in protein synthesis, he and his team pioneered methods for oligo synthesis that are still being used today. One of their many important contributions was the protection chemistry necessary for the sequential addition of a base nucleotide. Khorana introduced the dimethoxytrityl (DMT) group, which protects the 5’ hydroxyl and is selectively removed under acidic conditions to present the reactive site for the next phosphate linkage5. He and his team also introduced protecting groups for the nucleoside bases: benzoyl, acetyl, and isobutyl groups to protect the exocyclic amines that are otherwise reactive to the conditions of synthesis. These protecting groups are still used today in oligo synthesis.
While Khorana’s advancement in the chemical synthesis of DNA was an amazing feat, it also had limitations. The oligonucleotide had to be purified after each nucleotide addition to remove byproducts. In 1970, Robert Letsinger improved oligo synthesis further with the introduction of phosphotriester chemistry6, and introduced the solid phase support for the manufacture of oligonucleotides (Figure 1). Letsinger showed that an oligonucleotide could be synthesised while attached to polystyrene support and then cleaved from the support to release the desired oligonucleotide7. The use of the polystyrene support also allowed for unwanted byproducts from the nucleotide addition to be washed away. Using a solid support and the phosphotriester chemistry revolutionized the synthesis of DNA. Letsinger later introduced the oxidation of the phosphite (P(lll)) to the pentavalent phosphate(P(V)) using iodine and water in THF8. This is known as the phosphite triester chemistry. The iodine/water based oxidiser is still used today as the oxidising reagent in oligonucleotide synthesis.
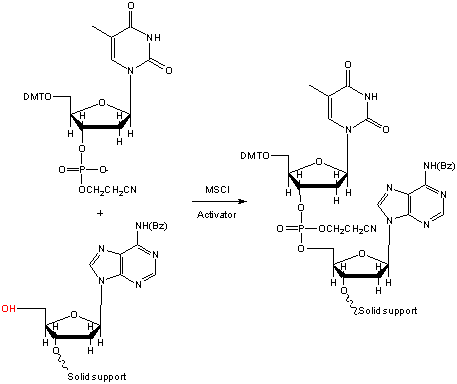
Figure 1. Phosphotriester chemistry
While the phosphotriester chemistry was the prominent chemistry used for DNA synthesis, it had disadvantages, such as long coupling times, unstable reagents, and low yields. In the early 1980’s, a student of Khorana and Letsinger, named Marvin Caruthers, introduced reagents that were much more efficient and stable in the form of cyanoethyl-protected phosphoramidite chemistry (Figure 2), revolutionizing DNA synthesis once again9.
Caruthers also introduced the use of controlled pore glass (CPG) as the support for the primary attachment of the growing oligonucleotide, which had advantages over the polystyrene support used previously10. Today, both types of supports are used in the chemical synthesis of DNA, while the phosphoramidite method is used on all synthesizers.
All of these advancements in nucleic acid chemistry ultimately led to the automation of oligonucleotide synthesis, with a microprocessor orchestrating the cycle of chemical reactions. In 1980, Vega Biosciences built the first DNA synthesizer, while Ron Cook of Biosearch developed the first commercially successful model instrumentation a couple of years later. Called the Synthesis Automation Machine, or SAM 1, this first modern synthesizer could make an abundance of oligonucleotides in a short amount of time, compared to the previous bench-top method of synthesis, and gave rise to the invention of PCR by Kary Mullis11.
Each of these forward thinking chemists contributed to the ability to synthesize the PCR primers, probes, and other tools used today. The chemistry has even advanced to the stage where it is possible to manufacture oligonucleotides that are 150-250 base pairs as well as large amounts of synthetic DNA.
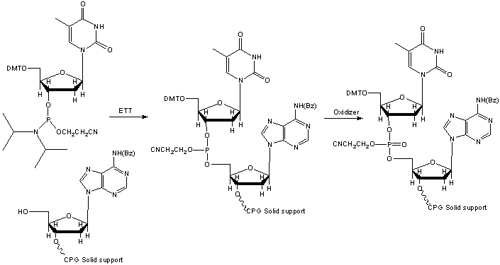
Figure 2. Solid Phase Phosphoramidite Coupling
The chemical synthesis of DNA
The principal reagents of the phosphoramidite method are the solid support such as a functionalized CPG, a phosphoramidite monomer, the detritylation reagent used to remove the DMT group, the activator, the capping reagent, and the oxidizing reagent.
The CPG is the solid support to which the first base is pre-attached, or derivatized, with a label to modify the 3’ position of the oligonucleotide. The solid support is packed into a column or reaction vessel for synthesis. As the reagents pass though the column, the oligonucleotide grows in the 3’ to 5’ direction, but remains attached to the CPG support throughout the synthesis.
The phosphoramidite monomer or modification is a nucleoside base, or else the reactive form of another modification that is added sequentially to the growing strand. The 5’ hydroxyl of the phosphoramidite is protected with a trityl group, such as DMT or MMT. A beta-cyanoethyl group protects the oxygen on the phosphormamidite and the reactive exocyclic amine groups of the base monomers are protected with conventional protecting groups. Some of these protecting groups were established in the phosphodiester chemistry from Khorana’s lab.
The synthesis cycle consists of four major steps (Shown in Figure 3), and culminates in the addition of a single residue to the growing oligo strand:
- The first step involves the removal of the trityl protecting group to generate the 5’ hydroxyl reactive site on the base modified CPG or base addition. A detritylation reagent, such as 3% dichloroacetic acid in dichloromethane, is used to remove the trityl protecting group and present the reactive site on the base-modified CPG. This leaves a 5’ hydroxyl group available for coupling during the next step of the synthesis.
- The second step in the cycle is the coupling step which involves activation of the phosphoramidite through the addition of tetrazole. As the activator and the phosphoramidite pass over the CPG, they form a tetrazolide intermediate. This intermediate reacts with the 5’ hydroxyl generated by the removal of the trityl group in the first step, and the formation of the 3’-5’ phosphite linkage to the growing oligonucleotide.
- The third step of the synthesis cycle is the capping of the unreacted 5’-hydroxyl sites with an acetyl group, to prevent the formation of heterogeneous oligonucleotide products. Capping of the unreacted 5’ hydroxyl sites ensures that only the complete chains of the oligonucleotide will continue growing.
- The fourth step is the oxidation of the phosphite triester to a more stable phosphate triester linkage, using an iodine solution. This step completes the cycle for the addition of a single base monomer or internal modification. The cycle is then repeated until the oligonucleotide is the desired length.
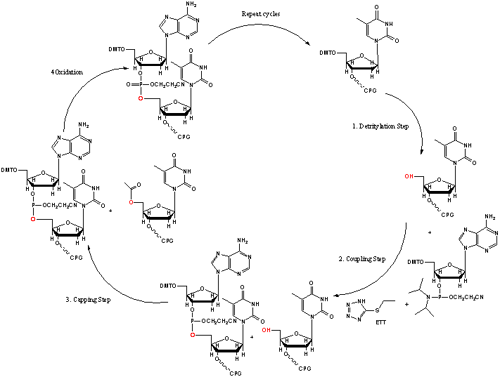
Figure 3. Oligo Synthesis Cycle
Once the desired sequence is synthesised, the oligo is exposed to basic conditions for the final work-up. This treatment serves to cleave the oligo from the 3’ support while simultaneously removing the protecting groups on the base monomers and phosphate linkages (cyanoethyl groups). After cleavage and deprotection, the final oligo can be purified and formulated for use in a genetic assay.
LGC Biosearch Technologies offers all standard base controlled pore glass supports, reagents, and dyes (DNA synthesis reagents) to chemically modify your oligo, as well as the knowledge and expertise in DNA synthesis chemistry since 1983 to synthesise your custom oligonucleotides.
References
- Nucleotides Part XXXll. Synthesis of a Dithymidine Dinucleotide Containing a 3’-5’- Internucleotide Linkage. Michelson, AM.; Todd, A. J .Chem Society. 1955; 0: 2632-38. doi: 10.1039/JR9550002632.
- Studies on Polynucleotides. I. A New and General Method for the Chemical Synthesis of the C5″-C3″ Internucleotidic Linkage. Syntheses of Deoxyribo-dinucleotides. Gilham, PT.; Khorana, HG. J Am Chem Soc. 1958 Dec; 80(23): 6212-22. doi: 10.1021/ja01556a016.
- Nobel Lecture: Nucleic Acid Synthesis in the Study of the Genetic Code. Khorana, HG. Nobel Lectures, Physiology or Medicine 1963-1970. 1968 Dec 12. Web. 1 Jul 2015. <http://www.nobelprize.org/nobel_prizes/medicine/laureates/1968/khorana-lecture.html>
- Total Synthesis of a Gene. Khorana, HG. Science. 1979 Feb 16; 203(4381): 614-25. doi: 10.1126/science.366749.
- Studies on Polynucleotides. XXIV The Stepwise Synthesis of Specific Deoxyribopolynucleoticles (4). Protected Derivatives of Deoxyribonucleosides and New Syntheses of Deoxyribonucleoside-3' Phosphates. Schaller, H.; Weimann, G.; Lerch, B.; Khorana, HG. J Am Chem Soc. 1963 Dec; 85(23): 3821-27. doi: 10.1021/ja00906a021.
- Nucleotide chemistry. XIII. Synthesis of oligothymidylates via phosphotriester intermediates. Letsinger, RL.; Ogilvie, KK. J Am Chem Soc. 1969 Jun; 91(12): 3350. doi:10.1021/ja01040a042.
- Oligonucleotide Synthesis on a Polymer Support. Letsinger, RL.; Mahadevan, V. J Am Chem Soc. 1965 Aug; 87(15): 3526-27. doi: 10.1021/ja01093a058.
- Nobel Lecture: Nucleic Acid Synthesis in the Study of the Genetic Code. Mullis, KB.; Smith, M. Nobel Lectures, Chemistry 1991-1995. 1993 Dec 8. Web. 1 Jul 2015. < http://www.nobelprize.org/nobel_prizes/chemistry/laureates/1993/mullis-lecture.html>
- Nucleotide Chemistry. XX. Phosphite Coupling Procedure for Generating Internucleotide Links. Letsinger, RL.; Finnan, JL.; Heavner, GA.; Lunsford, WB. J Am Chem Soc. 1975 May; 97(11): 3278-79. doi: 10.1021/ja00844a090.
- An investigation of several deoxynucleoside phosphoramidites useful for synthesizing deoxyoligonucleotides. McBride, LJ.; Caruthers, MH. Tetrahedron Letters. 1983; 24(3): 245-48. doi: 10.1016/S0040-4039(00)81376-3.
- Synthesis of deoxyoligonucleotides on a polymer support. Matteucci, MD.; Caruthers, MH. J Am Chem Soc. 1981 Jun; 103(11): 3185-91. doi: 10.1021/ja00401a041.

