Originally published : Fri, February 20, 2015 @ 5:52 PM
 Updated : Mon, November 16, 2015 @ 3:55 PM
Updated : Mon, November 16, 2015 @ 3:55 PM
Buffers are often overlooked and taken for granted until the day comes when a peculiar result is observed and its origin is traced to a bad buffer. Although in very rare cases mistakes in the composition of buffers have led to discoveries such as the correct number of human chromosomes1, using properly prepared buffers can be the key to success with technologies such as Stellaris RNA FISH.
Buffers are employed for any experiment that requires the structure and/or activity of biological material to be maintained. They function to resist changes in hydrogen ion concentrations and to provide critical salt concentrations, nutrients (such as for cell culture), and co-factors required for enzymatic reactions to take place. The new Stellaris RNA FISH Buffers come with 4 major advantages:
- Clarity: You may be thinking about diamonds here. Not quite! Single RNA molecules are clearer to identify because they are not masked by as much background fluorescence. The buffers contain proprietary additives to enhance detection, particularly for assays that exhibit more pronounced background fluorescence. We have observed, in over 50% of probe sets, pronounced background reduction effects compared to conventional RNA FISH buffers
- Consistency: Scientists will not have to deal with variable results from poor buffer composition. Your results will remain consistent from experiment to experiment.
- Convenience: The buffers provide a convenient workflow solution for Stellaris applications. You won’t have to worry about whether your lab-mate used up the last of the dextran sulfate and didn’t order more. You also won’t have to spend precious time mixing buffers.
- Confidence: Increased clarity, consistency, and convenience will give you confidence in your results. Stellaris RNA FISH buffers contribute to a more robust system for RNA detection and analysis. Please see the figure below for our direct comparison experiments.
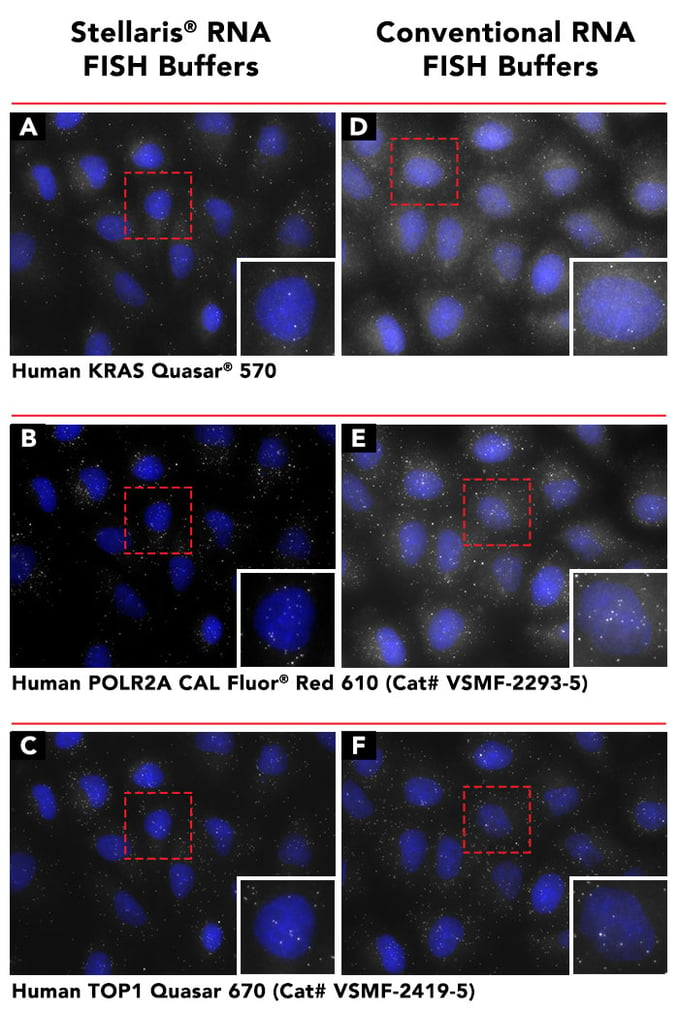
*View a hi-res version of the Stellaris FISH Buffers comparison
Fig - A direct comparison showing background reduction by utilizing Stellaris RNA FISH Buffers (Cat#: SMF-HB1-10, SMF-WA1-60, SMF-WB1-20). Following the adherent cell protocol, formalin fixed human cells were probed with Human KRAS, Human POLR2A (Cat# VSMF-2293-5) or Human TOP1 (Cat# VSMF-2419-5). Left Panel (A,B,C): RNA FISH performed with Stellaris RNA FISH Buffers. Right Panel (D,E,F): RNA FISH performed with Conventional RNA FISH buffer recipes. Note: Results may vary according to original probe set performance and sample type.
References:
- Mammalian Chromosomes In Vitro: I The Karyotype of Man J. Heredity. Hsu, T.C. 1952. 167-172

