Originally published : Wed, April 11, 2018 @ 2:25 PM
Updated : Mon, April 30, 2018 @ 1:53 PM
To truly understand complex diseases such as cancer, scientists need access to tools that don’t just average data across millions of cells, but that can interrogate single cells to examine gene expression and regulation in diseased cells.
While there are an increasing number of RNA FISH (fluorescence in-situ hybridisation) technologies available, LGC Biosearch Technologies‘ Stellaris® RNA FISH is the only technique that does not include signal amplification. This makes it a true orthogonal validation to RNA-seq or rtPCR and ensures results are truly quantifiable in a way that amplified signals can never be.
 Dr. Jessica Kaplunov, Product Marketing Manager at LGC Biosearch Technologies recently sat down with Biocompare to tell them more about this exciting technology and a number of new applications that are helping to make the world a healthier place.
Dr. Jessica Kaplunov, Product Marketing Manager at LGC Biosearch Technologies recently sat down with Biocompare to tell them more about this exciting technology and a number of new applications that are helping to make the world a healthier place.
According to Kaplunov, FISH analysis within a research setting is most commonly performed on cells grown in tissue culture or on fresh-frozen tissue samples, yet customers are continually discovering new applications for the technique. “We work closely with researchers to co-develop new applications and capabilities of RNA FISH, and there is considerable excitement and enthusiasm among thought leaders and within our group for approaches such as CASFISH, seqFISH and MERFISH, all of which hold possibilities for advancing the technology.”
“One major strength of RNA FISH is that it addresses the question of stochastic expression in a way that more classic techniques are unable to do,” says Kaplunov. “While we’ve known for a long time that expression of proteins and RNA between cells and even within a single cell can vary tremendously, techniques have routinely involved combining millions of cells together into one data point. Stellaris RNA FISH allows researchers to visualize expression patterns and target localization, and to quantify individual RNA molecules in situ to gain a much more nuanced understanding of expression and sequestering. Results generated with FISH add significant value to research, and this is exemplified by the exponential growth which we’ve seen in the number of papers citing Stellaris RNA FISH year on year, especially as a third of those are published in Nature, Cell or Science.”
Stellaris RNA FISH is quantifiable, capable of high order multiplexing, and offers exceptional specificity through a patented 48 probe technology. “This tiles up to 48 unique probes along the target RNA, each of which has a single small fluorophore, and only when dozens of probes are bound does the combined fluorescence become visible as a spot,” elaborates Kaplunov. “A Stellaris RNA FISH assay can be completed in a single day; furthermore, it can be combined with immunofluorescence to generate images of both RNA and proteins labeled together.”
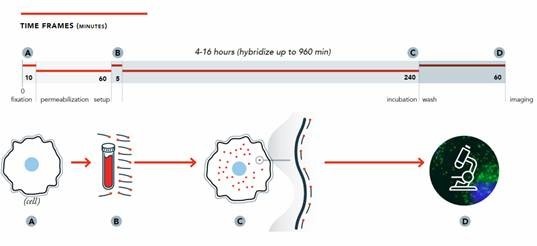
The Stellaris RNA FISH methodology
Kaplunov adds that she particularly likes the use of seqFISH to identify targets of interest followed by Stellaris RNA FISH assays to visualize, localize and quantify those targets in situ. “As discussed by Parker et al., Stellaris is a true orthogonal validation to RNA-seq.”.
“While qPCR is typically used as a validation tool for RNAseq, both techniques use primer-based amplification and could exhibit the same biases if any exist. Stellaris bridges the gap between more classic techniques and the stochastic expression and cell-specific data gained by FISH, and represents a logical next step in laboratory technique; providing a full range of data.”
To read the full Biocompare article head here.

