Originally published : Tue, October 26, 2021 @ 11:13 PM
Updated : Wed, December 14, 2022 @ 4:43 PM
This is the second blog post in a three-part educational series about best practices for in-house oligonucleotide synthesis.
In our first oligonucleotide synthesis basics blog post, we introduced phosphoramidite chemistry and discussed considerations for in-house and outsourced oligo synthesis. In this post, we continue to explore oligonucleotide synthesis and will examine key takeaways for each step of the synthesis cycle.
Deblocking (detritylation)
Starting with the support bound nucleoside or modifier, the first step of the oligonucleotide synthesis cycle is deblocking or detritylation. This involves removing the dimethoxytrityl protecting group from either the 5’ position of the nucleoside or from the modifier backbone with an acid treatment to afford a reactive OH group. The 4,4'-dimethoxytrityl (DMTr) protecting group is required, specifically on phosphoramidites, to prevent self coupling during the coupling step.
DMTr is commonly used for the protection of OH groups because it is easily removed with a dichloroacetic (DCA) or trichloroacetic acid (TCA). However, prolonged contact with acid can result in side reactions such as depurination, especially when assembling oligonucleotides that contain polyA sequences or sensitive nucleosides. Some examples of bases vulnerable to depurination include N(6)-benzoyl-2-deoxyadenosine, 8-aryl derivatives of 2-deoxyguanosine and to a lesser extent, N(2)-isobutyryl-2-deoxyguanosine.1
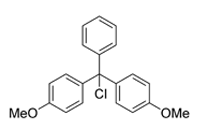
4,4-Dimethoxytrityl Chloride (DMTr-Cl), a reagent for
creation of reactive DMT functionality on nucleosides.
Depurination can lead to chain cleavages under basic conditions, such as during the final cleavage and deprotection steps to cleave the oligonucleotide from the support and remove the nucleobase and phosphorus protection. In some cases, using weaker acids during detritylation or reducing acid contact times can be beneficial to maintain low levels of depurination.1 DCA is a milder acid than TCA, making it a good option for detritylation of long oligonucleotides and tends to result in better synthetic yields than TCA.2 However, using TCA for deblocking allows for faster reactions. When 3% TCA was used in the detritylation step, studies show that acid delivery time as short as 10 seconds (versus 110 seconds) did not significantly compromise the yield of the full-length product.3 A deblocking step that completes in less than one minute is ideal for reducing the oligonucleotide’s exposure to acid.4
An alternative is to alternate the detritylation step with wash steps, this enables full deprotection but minimises the contact time with acid.
| Key takeaway: Use the mildest effective deblocking acid and shortest contact to reduce the risk of depurination. |
Activation/coupling
After the deblocking step is complete, the 5’-OH group is free for reaction. The phosphoramidite corresponding to the second base in the oligonucleotide sequence is activated. The diisopropylidene group is protonated by an activating agent such as 5-ethylthio-1H-tetrazole (ETT) or 5-benzylthio-1H-tetrazole (BTT). This is delivered to the column and reacts with the free 5’-OH group, displacing the diisopropylidene group and forming a phosphite linkage.
4,5-dicyanoimidazole (DCI) is an alternative activator that may be used. DCI is less acidic than ETT or BTT but is a more nucleophilic activator, reducing coupling times.5 It is highly soluble in acetonitrile and thus allows for higher effective concentrations of nucleoside phosphoramidites during solid-phase synthesis. A higher effective concentration, in turn, allows for lower phosphoramidite excess during coupling.5
Coupling efficiency is fundamental for optimising yield, oligonucleotide quality and for ease of the downstream purification process, especially for longer oligonucleotides. Many factors can affect the coupling efficiency of amidites including coupling time and acetonitrile dryness. In addition to using the most effective activator for the application, effective coupling efficiency can be achieved by:
- Eliminating moisture in reagents involved in the coupling step. Use anhydrous acetonitrile (ACN) as a diluent and wash and keep an anhydrous environment when dissolving phosphoramidites. We suggest pre-treating ACN with molecular sieves before adding it to the amidite.
 Considering oligo length, complexity and loading requirements when choosing a solid support. Starting with an appropriate solid support based on the oligonucleotide’s characteristics is critical to maintaining a high coupling efficiency throughout the synthesis cycle.
Considering oligo length, complexity and loading requirements when choosing a solid support. Starting with an appropriate solid support based on the oligonucleotide’s characteristics is critical to maintaining a high coupling efficiency throughout the synthesis cycle. - Increasing the phosphoramidite concentration to enhance the coupling efficiency for long oligonucleotides.2
- Adjusting reagent amounts and coupling time.
- For long oligos, using a cap/ox/cap cycle can improve coupling. This is thought to be a result of the second capping drying the support after the oxidation step.
Enhanced view of pore structure in controlled
pore glass (CPG) solid support.
| Key takeaway: Take steps to optimise coupling efficiency by optimising the coupling time and ensuring an anhydrous environment during the coupling process. |
Capping
Following phosphoramidite coupling, a capping step is introduced to acetylate any unreacted 5’-OH groups. While solid-phase phosphoramidite coupling usually proceeds to approximately 99% efficiency, any reactive 5’-OH groups left uncapped can result in undesirable side-products. Unless blocked, these truncated oligonucleotides can continue to react in subsequent synthesis cycles, creating near full-length oligonucleotides with internal deletions.

Capping is generally achieved using a solution containing acetic anhydride (Cap Mix A) and the catalyst N-methylimidazole (Cap Mix B), which acts as an activator for Cap Mix A. The 5′ acetyl ester cap is unreactive in all subsequent synthesis cycles and is removed during the final base deprotection step.4
Capping is a critical process in the synthesis cycle to reduce the accumulation of deletion mutations that are difficult to purify and could render the oligonucleotide ineffective for subsequent applications. Additional acetonitrile washing following capping may increase synthetic yield.4
Cap mix reagent bottles on a MerMade 192 synthesizer.
| Key takeaway: Even with the most efficient chemistry and purest reagents, it’s not reasonable to expect 100% coupling efficiency. |
Oxidation
In the final step of the oligonucleotide synthesis cycle, the unstable phosphite triester linkages are oxidised to a more stable pentavalent phosphotriester using iodine in a THF/water/pyridine or THF/water/lutidine solution. Because the oxidiser contains water, the support is rinsed several times with acetonitrile following this reaction.4 The most effective way to remove residual water is to perform an additional capping step. The pyridine or lutidine in the capping reagents remove the water from the support more efficiently than MeCN.
After oxidation is complete, the entire 4-step cycle is repeated, beginning with detritylation of the 5’-terminus of the support bound oligonucleotide. The process continues until each base of the oligonucleotide is coupled to the growing chain.
Our final blog post of this series covers post-synthesis processing. Stay tuned to learn about cleavage, deprotection and purification options for oligonucleotide synthesis.
| Key takeaway: While oxidation is the final step in the synthesis cycle, further processing of the oligonucleotide is required to make it a biologically active molecule. |
 |
If you are thinking about bringing oligo synthesis in-house, there are many considerations to account for beyond instrument selection. But the benefits of synthesising your own oligos are clear. Download this guide that summarises all the practical and technical information in a single place, to ensure that your synthesizers run at full capacity and you maximise your return on investment. |
References
- Tram K, Sanghvi Y and Yan H. Further optimization of detritylation in solid-phase oligodeoxyribonucleotide synthesis. Nucleosides, nucleotides & nucleic acids. 30:12-9. https://doi.org/10.1080/15257770.2010.537291. Published 2011. Accessed August 10, 2021.
- Pollard J. Synthesis of Oligonucleotides. Department of Molecular Biology - Massachusetts General Hospital and Harvard Medical School protocols. Published 1998. Accessed August 10, 2021. https://molbio.mgh.harvard.edu/szostakweb/protocols/oligo/index.html
- Millar S, Yan H, Sanghvi YS, et al. Some observations on detritylation in solid-phase oligonucleotide synthesis. Nucleic Acids Symp. Ser. 52(1):311–312. Published 2008. Accessed August 10, 2021. https://doi.org/10.1093/nass/nrn157.
- Ellington A and Pollard J. Introduction to the synthesis and purification of oligonucleotides. Current Protocols in Nucleic Acid Chemistry. A.3C.1-A.3C.22. Published 2000. Accessed August 10, 2021. https://deepblue.lib.umich.edu/bitstream/handle/2027.42/143751/cpnca03c.pdf?sequence=1
- Vargeese C, Carter J, Yegge J, et al. Efficient activation of nucleoside phosphoramidites with 4,5-dicyanoimidazole during oligonucleotide synthesis. Nucleic Acids Research. 26(4):1046-1050. https://doi.org/10.1093/nar/26.4.1046. Published 1998. Accessed August 10, 2021.

