Originally published : Wed, April 29, 2020 @ 3:17 PM
Updated : Wed, March 22, 2023 @ 1:52 PM
We’re not rejecting you, just your oligo
PCR is arguably one of the most enabling technologies of modern molecular biology, with a sensitivity that can detect incredibly small amounts of DNA. However, the sensitivity of the method is also one of the biggest challenges—when you can detect just a few molecules of DNA, inadvertent contamination can give you misleading data. Issues with contamination are especially critical in qPCR reactions for molecular diagnostics testing, such as for SARS-CoV-2 detection, where any contaminating template DNA, even if it’s just long fragments, can generate false positive results.
 The risks of contamination should be no surprise to any scientist that regularly uses PCR, but one consequence that you may not have considered is how this risk can affect your oligonucleotide order. Because oligos longer than 55 nt (a.k.a. longmers) are more likely to serve as templates for amplification, and because we recognize that there is a small but plausible risk of trace amounts of oligo from one order contaminating another order (via aerosolization), we have a very stringent process for screening orders for oligos longer than 55 nt. Any orders that could jeopardize the quality and reliability of any of our clients’ assays are rejected. In general, we do not synthesize any long positive control templates (unmodified oligos) and carefully screen other long modified oligos for their ability to contaminate other qPCR assays we sell.
The risks of contamination should be no surprise to any scientist that regularly uses PCR, but one consequence that you may not have considered is how this risk can affect your oligonucleotide order. Because oligos longer than 55 nt (a.k.a. longmers) are more likely to serve as templates for amplification, and because we recognize that there is a small but plausible risk of trace amounts of oligo from one order contaminating another order (via aerosolization), we have a very stringent process for screening orders for oligos longer than 55 nt. Any orders that could jeopardize the quality and reliability of any of our clients’ assays are rejected. In general, we do not synthesize any long positive control templates (unmodified oligos) and carefully screen other long modified oligos for their ability to contaminate other qPCR assays we sell.
The calculus behind rejection
The main factor we consider when we evaluate longmers is their potential to serve as a PCR amplification template. Thus, if your “longmer” has an internal dye, quencher or other polymerase blocking modification, it cannot serve as a template for amplification and we can synthesize it.
Similar considerations apply to what we call “functional” sequences, such as barcodes, NGS primer and adaptor sequences, and KASP motifs. We only consider the length of the oligo outside of these motifs and will synthesize an oligo that is, say 25 nt KASP motif and 25 nt target sequence.
Additionally, we can synthesize long oligo aptamers. These oligos do not serve as amplification templates and do not put other qPCR assays at risk.
We run additional analyses including BLAST searches and screen against recent oligo orders to assess the risk of any longmer to other assays we sell and only synthesize low risk longmers to ensure the quality of our customers’ assays.
However, there are certain high risk sequences we will not synthesize, such as any oligo or series of oligo fragments that could serve as a template for SARS-CoV-2:
To ensure continued delivery of quality, uncompromised oligos, we will not synthesize DNA or RNA templates or long oligos with the potential to be templates for SARS-CoV-2. Please understand this approach is taken strictly to protect the integrity of your COVID-19 lab work.
While this means that we will not synthesize positive control oligos for SARS-CoV-2 testing, we do offer robust, full-process quality solutions through LGC’s recent acquisition of SeraCare, which makes available both the AccuPlex™ SARS-CoV-2 Verification Panel and the AccuPlex™ SARS-CoV-2 Reference Material Kit. Note that these positive control materials are manufactured in a completely separate facility from our oligonucleotides—they are over 3,000 miles apart—and do not pose a contamination risk to your oligo order.
Longmers are not the only possible source of contamination
Of course, longmers are just one possible source of contamination in a PCR reaction, although the team at Biosearch Technologies works hard to keep this risk as low as possible in our facility. While we cannot be certain when or how a false positive emerged, we find that carry-over contamination in the lab performing the PCR reaction is the usual culprit, with aerosolization of the PCR product the most likely source.
Excellent outcomes notwithstanding, it’s possible for the laboratory atmosphere itself to become contaminated to the extent that the ambient environment is the common point of contact for the unexpected template (Figure 1). In this situation, the only remedy is to prepare the reactions within another area entirely if possible, never introducing the oligo stocks, ancillary reagents, or consumables into the suspect environment. In addition, we recommend that the reaction preparation area is carefully cleaned with a 10% bleach solution, including all surfaces and pipettes, and obtaining new aliquots of consumables where possible. Hopefully this will reduce the risk of contamination. You could also review this FAQ for additional suggestions to prevent contamination.
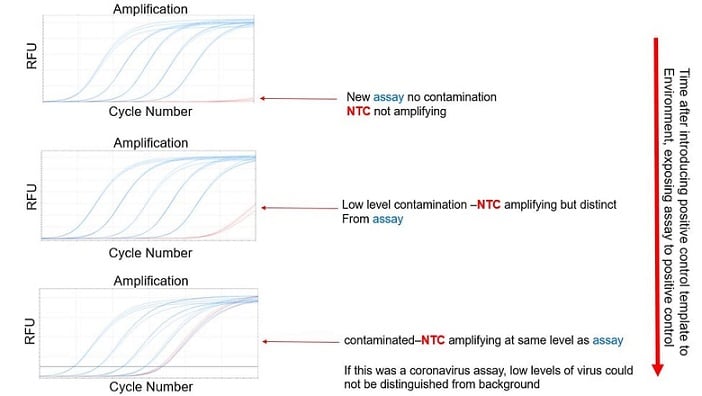
Figure 1. What contamination from an environmental source might look like.
The bottom line: We anticipate and mitigate risks to your studies
At Biosearch Technologies, we are serious about using science for a safer world. When it comes to our mission of providing critical reagents for molecular diagnostics, such as oligos, we work hard to ensure that you can have complete confidence in the products and services we deliver. Even if it means that sometimes, we just have to say no.
Is contamination impacting the quality and reliability of your qPCR assays?
Get specific recommendations to evaluate symptoms and diagnose the cause of your unexpected results in our white paper.
 |
If you are thinking about bringing oligo synthesis in-house, there are many considerations to account for beyond instrument selection. But the benefits of synthesizing your own oligos are clear. Download this guide that summarises all the practical and technical information in a single place, to ensure that your synthesizers run at full capacity and you maximise your return on investment. |


