Originally published : Mon, February 11, 2019 @ 5:12 PM
Updated : Fri, May 19, 2023 @ 8:53 AM
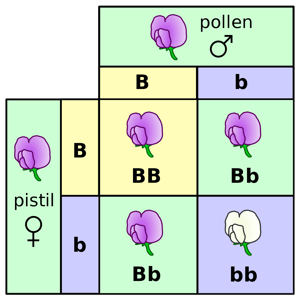
“Experience of artificial fertilisation, such as is effected with ornamental plants in order to obtain new variations in colour, has led to the experiments which will here be discussed. The striking regularity with which the same hybrid forms always reappeared whenever fertilization took place between the same species induced further experiments to be undertaken.”1
 Thus begins Gregor Mendel’s paper, published in 1866, one of the first documented examples of using genetics to develop and enhance varieties of plant crops. He was eventually recognised after his death and is often referred to as “the father of modern genetics”.
Thus begins Gregor Mendel’s paper, published in 1866, one of the first documented examples of using genetics to develop and enhance varieties of plant crops. He was eventually recognised after his death and is often referred to as “the father of modern genetics”.
Fast forward past the discovery of DNA and the genetic code, and we reach the exciting development of next generation sequencing (NGS) which is now a technology of choice used to develop higher yield crop varieties that can better tolerate drought and disease.
NGS is revolutionising agriculture, and genomic selection has become a realistic option for our future food supply.
The first completed version of the bread wheat genome sequence was assembled and published in 2018.2 It took 13 years and was a huge scientific challenge due to its sheer size: over 5 times bigger than the human genome, with over 80% repeated elements.
The human genome has about 3 billion base pairs, where bread wheat has around 17 billion, so without a technology such as NGS, this breakthrough would have been tough.
Curtis Pozniak,3 a leader in the International Wheat Genome Sequencing Consortium (IWGSC) uses NGS extensively and and drove the project to sequence 10 additional wheat genomes from various geographies. This revealed extensive structural rearrangements and difference in gene content resulting from complex breeding histories in the diverse environments.4
How does NGS for sequence-based genotyping enable scientists to reap the benefits of genomic selection?
NGS makes it much easier to apply genome selection, since breeders can more easily design and implement their breeding programmes to develop desirable traits such as drought tolerance, disease resistance, and higher yields. NGS enables genotyping by sequencing (GBS) and targeted genotyping by sequencing.
Looking at experimental design in this area, single nucleotide polymorphism (SNP) discovery involves 10,000-100,000 markers on perhaps as few as 5 samples, whereas the sweet spot for genomic selection is around 1,000-25,000 markers run on around 1,000 samples.
If one can apply such range of multiplexing using the same technology, it is obviously more efficient. NGS is that flexible technology, allowing one to use targeted genotyping by sequencing methods that can not only screen up to 100,000 markers per sample but also function in that mid-plex sweet spot of 500 to 25,000 markers.
Equally NGS can be used for whole genome sequencing, although in most crops the exome corresponds to only 1-2% of the entire genome, so to be cost effective, it is probably better to focus on the most relevant parts of the genome.
How does NGS-based SNP analysis compare with arrays?
 Arrays used to be considered the standard for SNP analysis, but by definition those arrays will only include existing, known SNPs. However, with sequencing-based methods such as targeted genotyping by sequencing, one can find new SNPs and structural variants in flanking sequences of targeted SNPs, and thus easily obtain much more genetic information.
Arrays used to be considered the standard for SNP analysis, but by definition those arrays will only include existing, known SNPs. However, with sequencing-based methods such as targeted genotyping by sequencing, one can find new SNPs and structural variants in flanking sequences of targeted SNPs, and thus easily obtain much more genetic information.
A study of 500 markers using sequences previously tested on an array, demonstrated that only 491 SNP sequences were originally selected to be common between the targeted genotyping by sequencing library and array data, whereas targeted genotyping by sequencing discovered 5,733 de novo SNPs.5
As a result, targeted genotyping by sequencing is starting to displace arrays for routine screening in breeding programmes. In the research labs at LGC Biosearch Technologies™ a wet lab targeted genotyping by sequencing experiment can take less than two weeks. If one adds several weeks to design a new oligo library, this means that a whole targeted genotyping by sequencing experiment can be fitted into a single plant breeding cycle, saving years of development time, essential as we tread the rocky road towards global food insecurity.
Read more about NGS in our post on NGS solutions for polyploid plant breeding challenges and our post on rice improvement.
At Biosearch Technologies we understand what real-world breeding programs need and we have bridged the gap in mid-plex genotyping technologies in developing our NGS-based targeted genotyping by sequencing, called Flex-Seq™. The Flex-Seq service has been shown to work for genome wide association studies (GWAS), marker-assisted selection/breeding (MAS/MAB), marker-assisted back crossing (MABC), QTL screening and trait mapping.
Find out more about Flex-Seq, our NGS-based targeted genotyping by sequencing service to help scientists make their genome selection projects a reality.
Our Flex-Seq service enables cost-effective screening for tens of thousands of markers. Flex-Seq uses multiplex hybridisation for cleaner results delivering up to 98% marker recovery. Customisable panels to which markers can be easily added or removed allow cost-effective flexibility versus chip-based arrays. The low cost per sample facilitates more screening, increasing yield and genetic gain and getting complex traits to market sooner.
References
- Mendel, G. (1866). Versuche über Plflanzen-hybriden. Verhandlungen des naturforschenden Ver-eines in Brünn, Bd. IV für das Jahr 1865, Abhand-lungen, 3–47.
- International Wheat Genome Sequencing Consortium (IWGSC, 2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome., Science, Vol. 361, Issue 6403.
- Curtis Pozniak (2015) http://www.wheatgenome.org/People/Leader-Spotlight/Curtis-Pozniak accessed 5 December 2018
- Walkowiak, S., Gao, L., Monat, C. et al. (2020). Multiple wheat genomes reveal global variation in modern breeding. Nature 588, 277–283. DOI:10.1038/s41586-020-2961-x
- Efficient genome-wide genotyping strategies and data integration in crop plants. (2018). Torkamaneh D et al. Theor Appl Genet. Mar;131(3):499–511.


